
Test on recursive parameter estimates, which are there? R-structchange also has musum (moving cumulative sum tests) SupLM, expLM, aveLM (Andrews, Andrews/Ploberger) Problems it should be also quite efficient as expanding OLS function. However, since it uses recursive updating and does not estimate separate This isĬurrently mainly helper function for recursive residual based tests. Test for model stability, breaks in parameters for ols, Hansen 1992Ĭalculate recursive ols with residuals and cusum test statistic. 
Unknown Change Point ¶ breaks_cusumolsresidĬusum test for parameter stability based on ols residuals Predictive test: Greene, number of observations in subsample is smaller than Known Change Point ¶ OneWayLS :įlexible ols wrapper for testing identical regression coefficients across Test whether all or some regression coefficient are constant over theĮntire data sample. Tests for Structural Change, Parameter Stability ¶ White’s two-moment specification test with null hypothesis of homoscedastic This tests against specific functional alternatives. Lagrange Multiplier test for Null hypothesis that linear specification isĬorrect. Multiplier test for Null hypothesis that linear specification is Non-Linearity Tests ¶ linear_harvey_collier Ljung-Box test for no autocorrelation of residualsīreusch-Pagan test for no autocorrelation of residuals durbin_watsonĭurbin-Watson test for no autocorrelation of residuals They assume that observations are ordered by time. This group of test whether the regression residuals are not autocorrelated. Test whether variance is the same in 2 subsamples Autocorrelation Tests ¶ Lagrange Multiplier Heteroscedasticity Test by White het_goldfeldquandt Lagrange Multiplier Heteroscedasticity Test by Breusch-Pagan het_white In the power of the test for different types of heteroscedasticity. Of heteroscedasticity is considered as alternative hypothesis. Heteroscedasticity Tests ¶įor these test the null hypothesis is that all observations have the sameĮrror variance, i.e. The following briefly summarizes specification and diagnostics tests for The second approach is to test whether our sample is To use robust methods, for example robust regression or robust covariance One solution to the problem of uncertainty about the correct specification is Only correct of our assumptions hold (at least approximately). Since our results depend on these statistical assumptions, the results are The errors are normally distributed or that we have a large sample. Homoscedasticity are assumed, some test statistics additionally assume that For example when using ols, then linearity and In many cases of statistical analysis, we are not sure whether our statistical If you are interested, see: How to Run and Interpret a Logistic Regression Model in R.Regression Diagnostics and Specification Tests ¶ Introduction ¶ Residual deviance: 116.96 on 98 degrees of freedom Null deviance: 138.63 on 99 degrees of freedom (Dispersion parameter for binomial family taken to be 1) Glm(formula = y_binary ~ x, family = "binomial") Summary(glm(y_binary ~ x, family = 'binomial')) # estimating heteroscedasticity-robust standard errors Here’s how to do it in R: model = lm(y ~ x) Since heteroscedasticity only biases standard errors (and not regression coefficients), we can replace them with ones that are robust to heteroscedasticity. Solution #2: Calculating heteroscedasticity-robust standard errors The log transformation did not solve our problem in this case since the residuals vs fitted values plot is still showing a fan shape instead of a random pattern. # re-check residuals vs fitted values plot The difference is that with a log transformation the model will still be interpretable, so this is what we are going to use here (for information on how to interpret such model, see: Interpret Log Transformations in Linear Regression): model1 = lm(log(y) ~ x) The first solution we can try is to transform the outcome Y by using a log or a square root transformation. 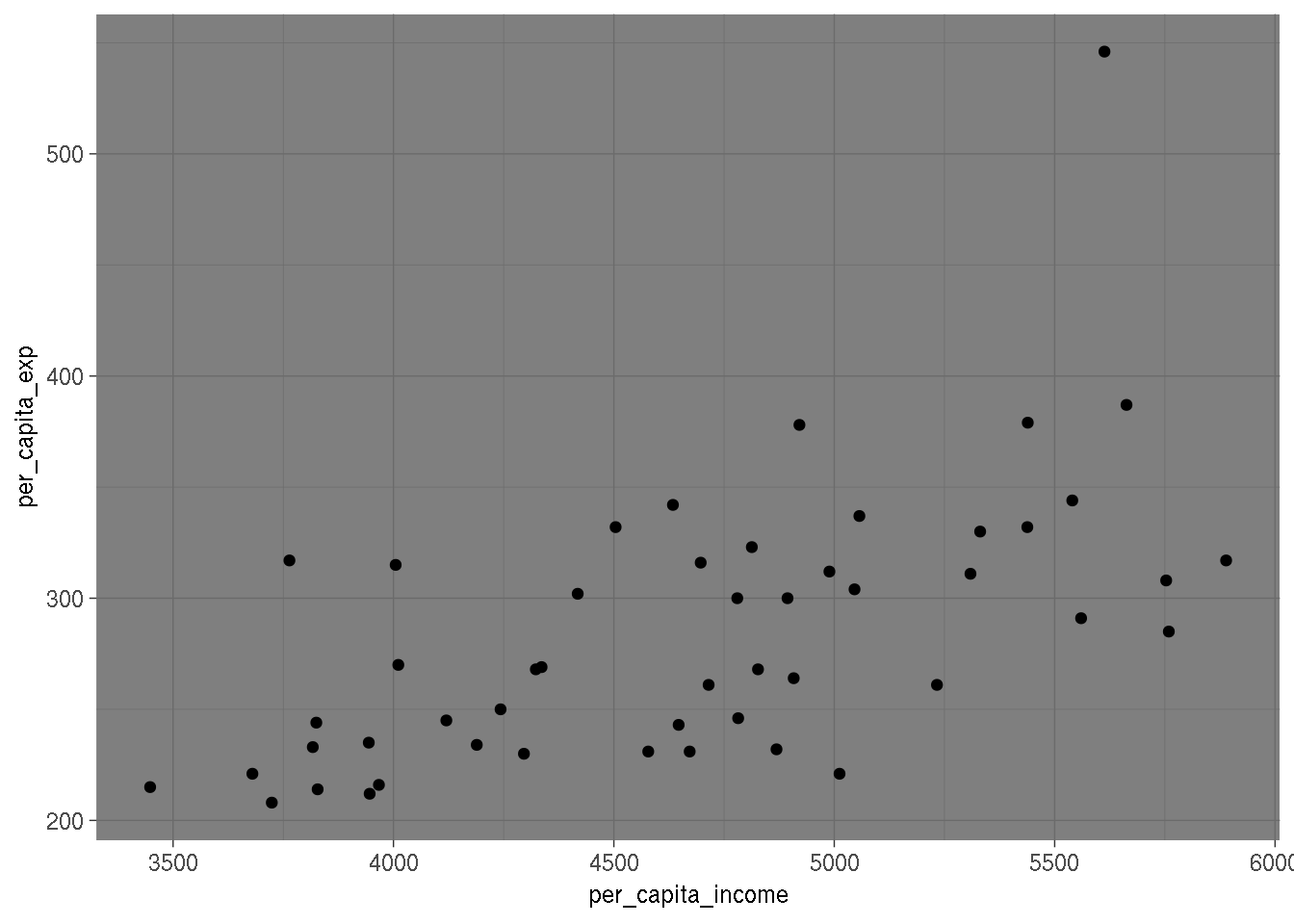
(For more information, see: How to Check Linear Regression Assumptions in R) Solution #1: Transforming the outcome variable The residuals vs fitted values plot shows a fan shape, which is evidence of heteroscedasticity. Let’s create some heteroscedastic data to demonstrate these methods: set.seed(0)
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
